Öffnet in neuem Fenster
Opens in a new window
Öffnet externe Seite
Opens an external site
Öffnet externe Seite in neuem Fenster
Opens an external site in a new window
James McNally
Biological Applications
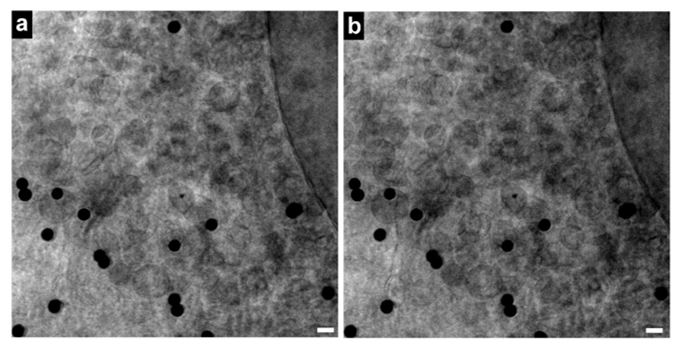
As long as specimens are cryo-preserved, X-ray imaging does not disrupt cellular structures. Shown below is a specimen imaged before and after the tomographic tilt series. Current resolutions are on the order of ~50 nm in xy and ~100 nm in z. These resolutions are currently set by limitations in the nanofabrication of the X-ray zone plate objectives and by limitations in the reconstruction procedures for the tomographic data.
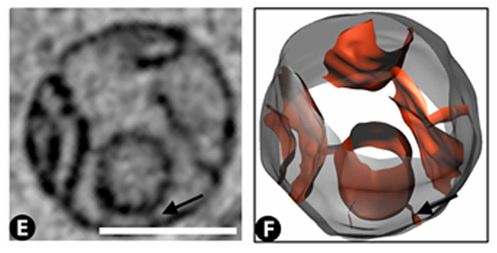
Slice through a mitochondria, and rendering of the mitochondria obtained from segmentation of the 3D X-ray image. Scale bar = 500 nm.
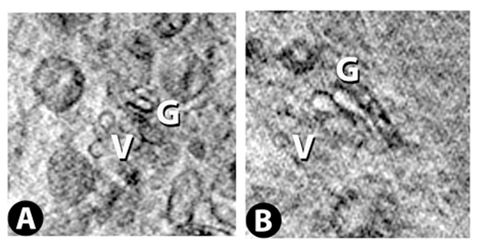
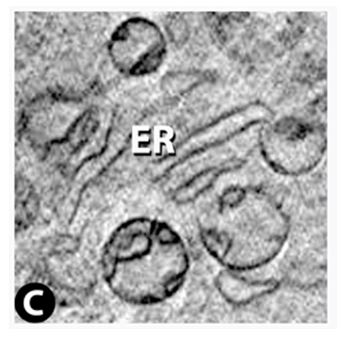
Two successive slices through the Golgi apparatus. Virtual slice thickness is ~30 nm.
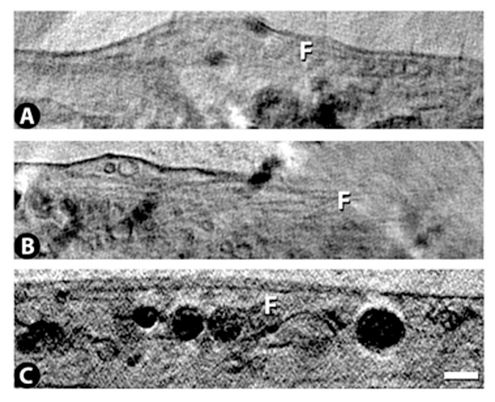
Filaments, presumably microtubules or microfilaments. Scale bar = 500 nm.
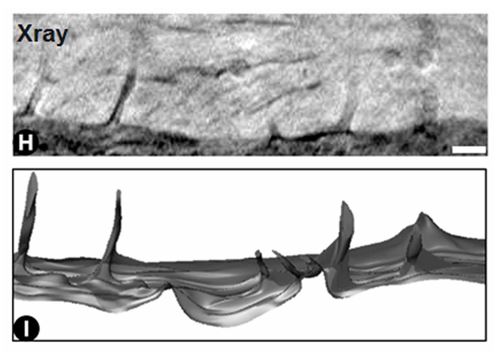
A slice through filopodia, and a 3D rendering obtained from segmentation of the 3D X-ray image. Scalebar = 500 nm.
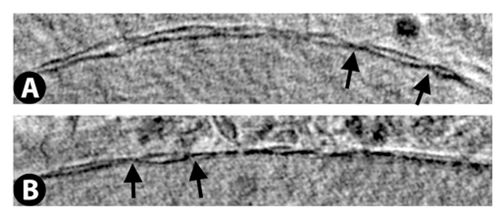
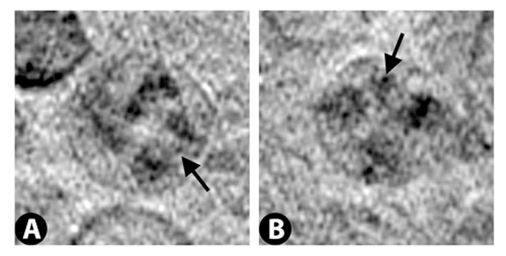
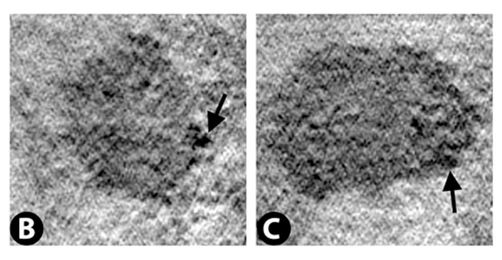
Nuclear membrane with nuclear pores indicated by arrows.
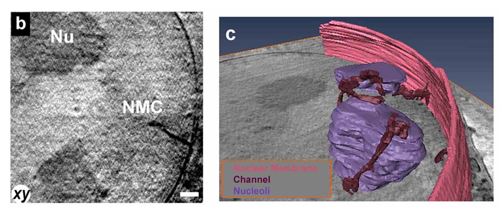

Nuclear membrane channels. Invaginations of the nuclear membrane often terminating at nucleoli. 3D rendering obtained from segmentation of the X-ray data. Scale bar = 390 nm.