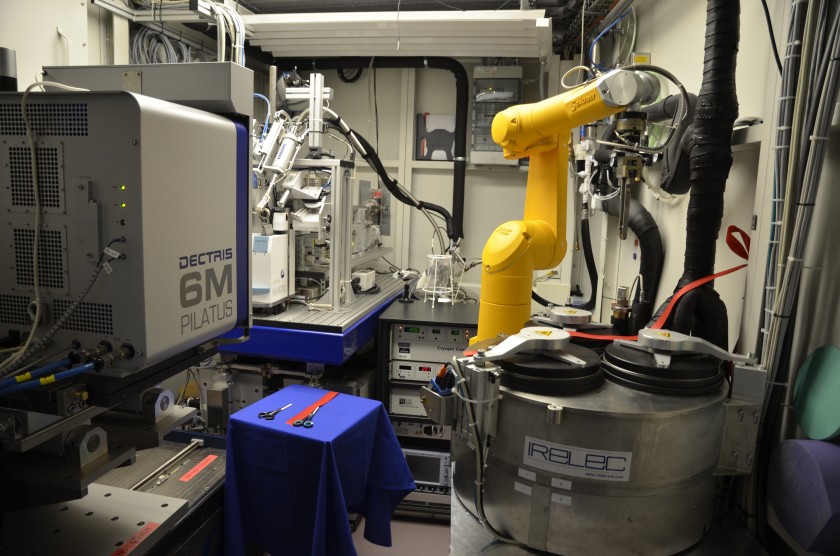
Protein crystallography at BESSY II: a tool for decoding the building blocks of life
Over 4.000 structures have been decoded at BESSY II (as of Dezember 2022), here you can get a picture of them
BESSY II is a brilliant light source for research, with soft and tender X-rays. With its special synchrotron light, we study proteins that play a crucial role in the functioning of viruses, bacteria, and cells, for example.
Protein molecules are involved in nearly all biochemical processes of living organisms and viruses. A single macromolecule can be made up of hundreds of amino acids, forming a huge and highly complex three-dimensional architecture. Parts of these macromolecules are folded, other parts look like loops or spirals, and there can be branches and symmetries. This 3D architecture determines how a protein functions.
How does protein crystallography work?
There are three beamlines dedicated to protein crystallography at BESSY II. These beamlines are also called the MX beamlines. The abbreviation MX stands for “Macromolecular X-ray Structure Analysis”. They can be used to decode the 3D architecture of proteins.
In the first step, proteins must be crystallised out of a protein solution so that the macromolecules arrange themselves into geometrically regular lattice sites. Growing crystals from proteins requires a lot of experience. Protein crystals are tiny and delicate. A team from the MX group at HZB has developed a special sample holder to considerably simplify the handling of these delicate crystals.
In the second step, the samples are cooled in liquid nitrogen, then gradually scanned in the X-ray beam using a robotic arm. The X-ray light is diffracted by the protein crystal structure, creating a diffraction pattern in the detector.
In the final step, each individual spot of this diffraction pattern is analysed. The 3D architecture of the protein can be decoded with the help of sophisticated algorithms. In addition, a computer programme also calculates the electron density around the protein molecule that contains important information about binding abilities and chemical properties of the protein.
The MX team at BESSY II
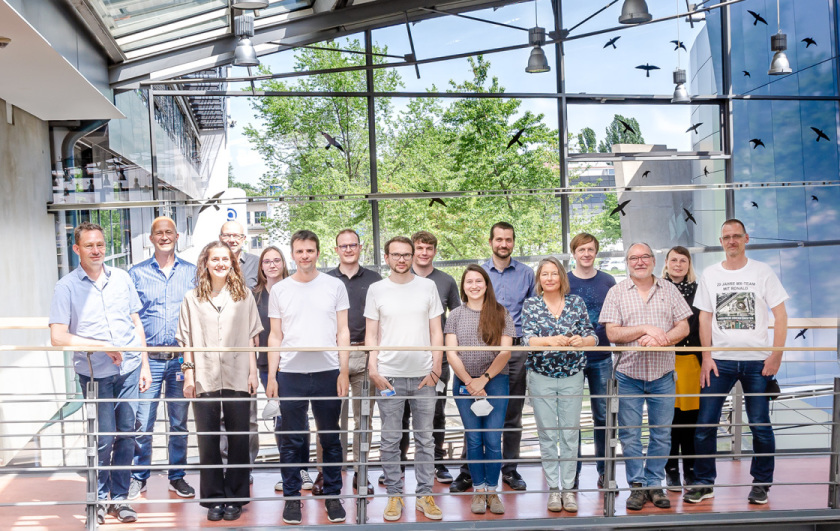

The Joint Research Group Macromolecular Crystallography, or MX team for short, has grown a lot over the years. It now numbers 12 people as of spring 2021: one PhD student, three engineers, a lab assistant, a research assistant, two postdocs, and four scientists.
Work at the beamlines during the pandemic looks different than usual. Due to the remote-work regulations that mandate single on-site staffing, the team often has to coordinate who works on site and who works from home. Doing research and lab work from home is complicated. Often, the only thing that can be done is processing the data and writing publications. While some staff still must be on site to keep the beamlines operating, hardly any external users can visit.
Instead, the first MX beamline runs completely via remote access. This means the users send in their samples to us, the team prepares them for measurement, and the rest is done by the users in front of their computers. They can control (almost) everything from there. The second beamline is presently being prepared for remote access. The third one has no automated sample changer and is therefore not suitable for remote operation. The SARS-CoV-2 virus (> article about the research) is an important research topic for the MX team; fragment screening is also an increasingly important activity. The MX team has attracted funding for this work from the EU and the Helmholtz Initiative and Networking Fund.

