Joint Research Group Macromolecular Crystallography
Structure of the month - January 2009
Structure Vol. 16, May 2008, Pages 766-77
An Intersubunit Active Site between Supercoiled Parallel β Helices in the Trimeric Tailspike Endorhamnosidase of Shigella flexneri Phage Sf6
Jürgen J. Müller1,4, Stefanie Barbirz2,4, Karolin Heinle2, Alexander Freiberg2,5, Robert Seckler2, and Udo Heinemann1,3,*
1 Max-Delbrück-Centrum für Molekulare Medizin, Robert-Rössle-Str. 10, 13125 Berlin, Germany
2 Physikalische Biochemie, Universität Potsdam, Karl-Liebknecht-Str. 24-25, 14476 Golm, Germany
3 Institut für Chemie und Biochemie, Freie Universität, Takustr. 6, 14195 Berlin, Germany
4 These authors contributed equally to this work.
5 Present address: The University of Texas Medical Branch, Department of Pathology, Keiller Building, 3.144, 301 University Boulevard, Galveston, TX 77555-0609, USA.
*Correspondence: heinemann@mdc-berlin.de
Abstract
Sf6 belongs to the Podoviridae family of temperate bacteriophages that infect gram negative bacteria by insertion of their double-stranded DNA. They attach to their hosts specifically via their tailspike proteins.
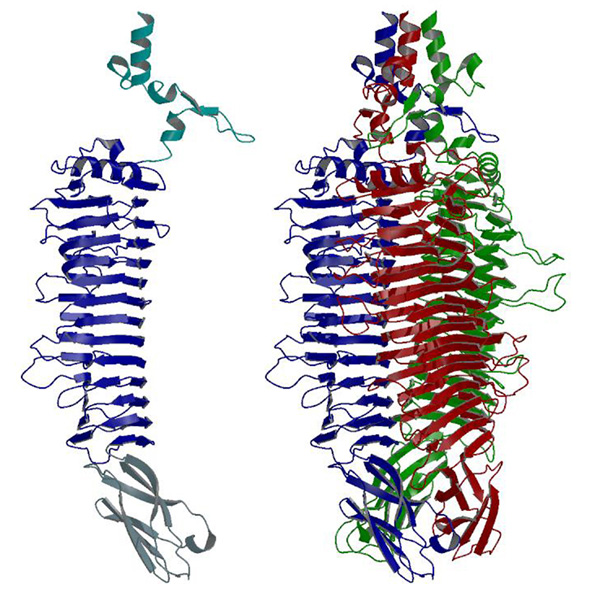
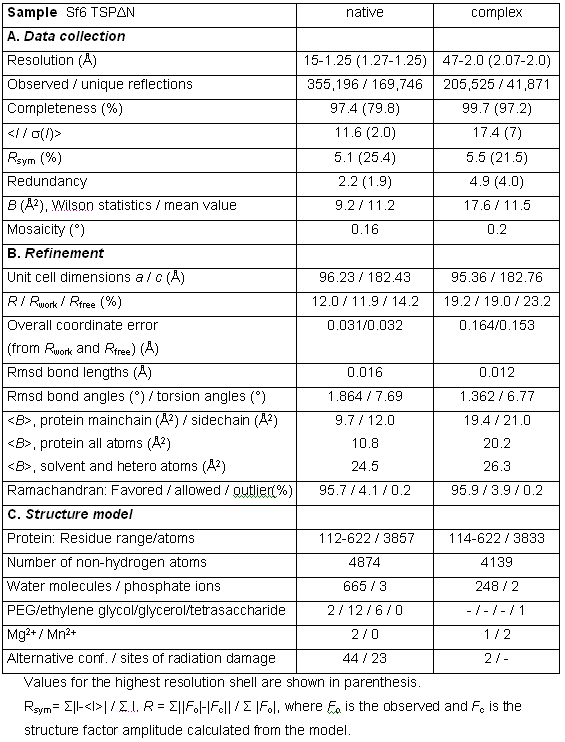
The structure of Shigella phage Sf6 tailspike protein lacking the head binding domain (Sf6 TSPΔN) was determined to a resolution of 1.25 Å at the EMBL BW7B beamline of DESY (Hamburg), whereas the diffraction data of the Sf6 TSPΔN in complex with the oligosaccharide were measured at the Protein Structure Factory Beamline BL 14.2 at BESSY (Table 1). The final model shows one monomer in the asymmetric unit. The biologically active trimer is built by crystallographic three-fold symmetry.
The monomer structure reveals a conserved architecture with a central, right-handed β helix. In the trimer of Sf6 TSPΔN, the parallel β helices form a left-handed, coiled β coil with a pitch of 340 Å. The C-terminal domain consists of a β sandwich reminiscent of viral capsid proteins. Further crystallographic and biochemical analyses show a Shigella cell wall O-antigen fragment to bind to an endorhamnosidase active site located between two β helix subunits each anchoring one catalytic carboxylate. The functionally and structurally related bacteriophage, P22TSPΔN, lacks sequence identity with Sf6 TSPΔN and has its active sites on single subunits. Sf6 TSP may serve as an example for the evolution of different host specificities on a similar general architecture.
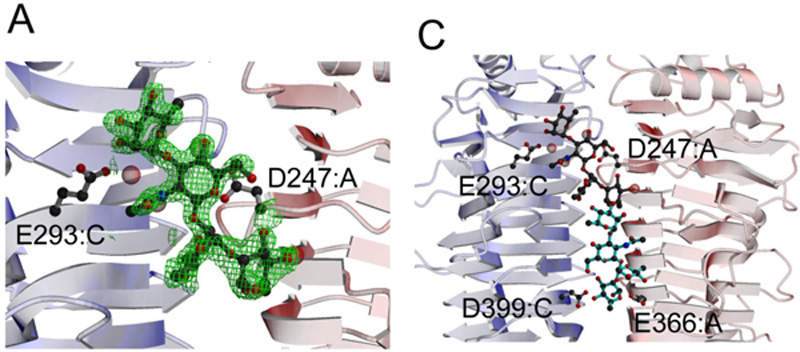
Figure 2. Localization of the Endorhamnosidase Active Site of Sf6 TSPΔN (PDB code 2VBM).
(A) Difference electron density (contoured at 3σ) observed in a complex of the protein with one repeating unit (RU) of an O-antigen hydrolysis product (a-L-Rhap-(1-3)-b-L-GlcpNAc -(1-2)-a-L-Rhap-(1-2)-a-L-Rhap). Glu293 (chain C) and Asp247 (chain A) belong to the binding site.
(C) An octasaccharide (2 RU) modeled into the binding site with its reducing end reaching the catalytic residues Asp399 and Glu366, which lie on different chains, as indicated (right). Bridging water molecules are colored purple.