Joint Research Group Macromolecular Crystallography
Structure of the month - August 2005
Nature Vol. 436, 2005, Pages 979-984
The structure of an bacterial ADP-ribosylating toxin in complex to its intracellular substrate
René Jørgensen1, A. Rod Merrill2, Susan P. Yates2, Victor E. Marquez4, Adrian L. Schwan3, Thomas Boesen1, and Gregers R. Andersen1
1 Centre for Structural Biology, Department of Molecular Biology, University od Aarhus, Gustav Wieds Vej 10C, DK-8000, Denmark
2 Department of Molecular and Cellular Biology, and
3 Department of Chemistry, University of Guelph, Ontario N1G 2W1, Canada
4 Laboratory of Medical Chemistry, Center for Cancer Research, National Cancer Institute at Frederick, NIH, Frederick, Maryland 21702, USA
Abstract
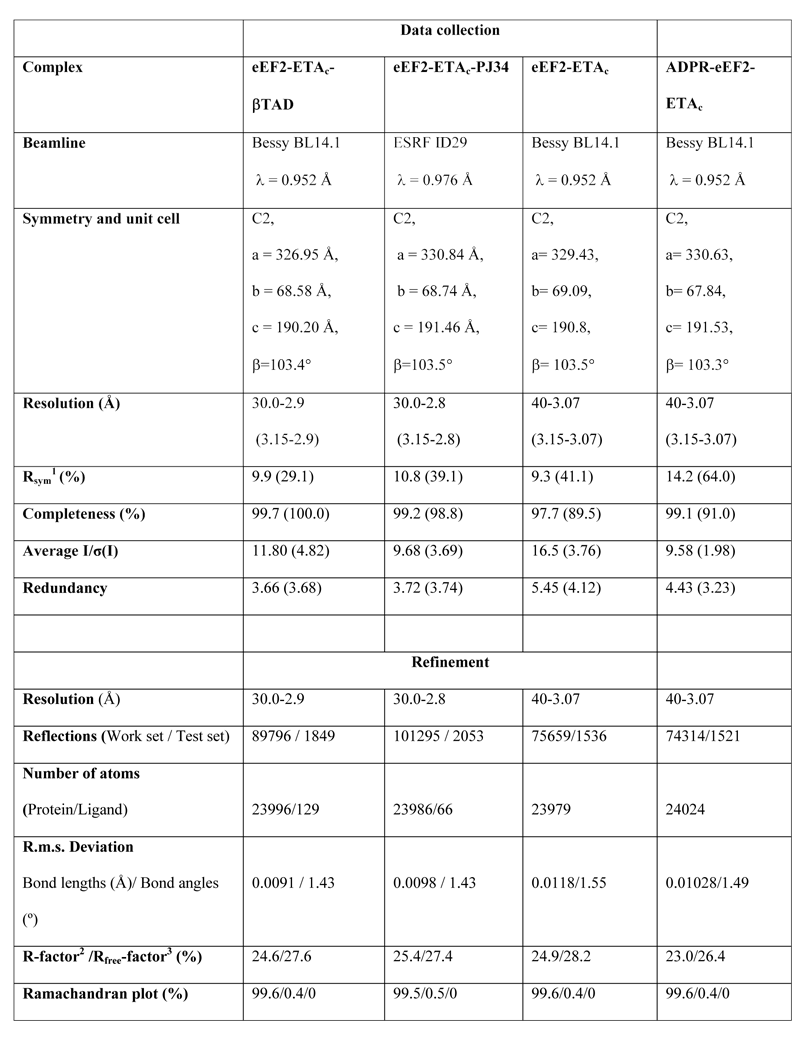
In a new Nature paper, researchers at University of Aarhus, Denmark, have now provided a structural basis for dissecting the enzymatic mechanism of ADP-ribosylating toxins, which comprises among others cholera, pertussis, diphtheria, and Pseudomonas exotoxin A. These toxins transfer an ADP-ribosyl group from NAD to a specific side chain of the target causing their inactivation. The team has determined the crystal structure of Pseudomonas exotoxin A in complex with its intracellular target translation elongation factor 2 (eEF2), which is ADP-ribosylated on the unusual aminoacid diphthamide (Figure 1). Elongation factor 2 is also the target of diphtheria toxin. ADP-ribosylation of elongation factor 2 by diphtheria toxin and exotoxin A leads to shutdown of translation and cell death. Four structures representing various steps in the enzymatic reaction were determined, and three of these were based on x-ray diffraction data collected at BESSY BL1 (table 1). In the crystal structure of the Michaelis complex, the distance between the electrophile and the nucleophile is 11 Å, which must be reduced below 3 Å at the transition state, but the mechanism for reducing this distance is still unknown. However, the enzymatic reaction takes place in the crystalline state proving the physiological relevance of the structures.
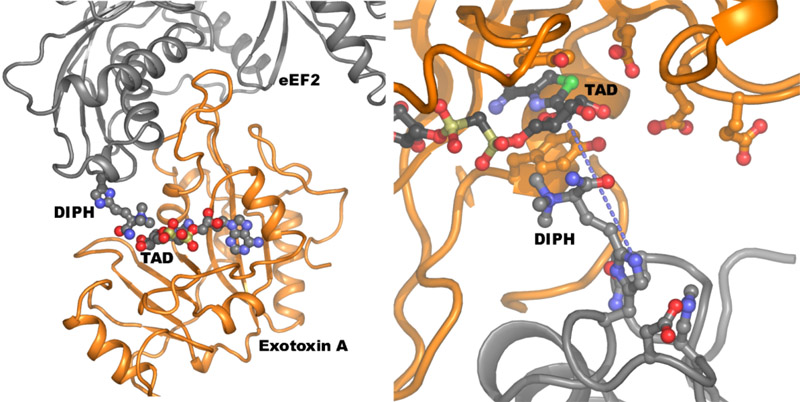
Figure 1. Left: The crystal structure of exotoxin A (orange) bound to its target eEF2 (grey), which is inactivated by the ADP-ribosylation transferred to the diphthamide residue (labelled DIPH). The TAD molecule is a nonhydrolyzable NAD analogue. Right: Close-up on the active site of the toxin enzyme in the Michaelis complex. The blue dotted line connects the two atoms to be connected at the end of the reaction.