Joint Research Group Macromolecular Crystallography
Structure of the month - December 2010
Mol. Cell Vol. 40, 658-670 (2010).
Crystal structures of RNase H2 in complex with nucleic acid reveal the mechanism of RNA-DNA junction recognition and cleavage
Monika P. Rychlik1, Hyongi Chon2, Susana M. Cerritelli2, Paulina Klimek1, Robert J. Crouch2 & Marcin Nowotny1*
1 Laboratory of Protein Structure, International Institute of Molecular and Cell Biology, 4 Trojdena Street, 02-109, Warsaw, Poland
2 Program in Genomics of Differentiation, Eunice Kennedy Shriver, National Institute of Child Health and Human Development National Institutes of Health, Bethesda, MD 20892, USA
* email: mnowotny@iimcb.gov.pl
Abstract
RNases H are divided into two main classes called type 1 and 2. RNase H1 in mice is important for mitochondrial DNA amplification and mutations in human RNase H2 cause Aicardi-Goutières syndrome, a severe neurological disorder. The preferred substrate of type 1 enzymes is an RNA/DNA hybrid with one strand containing a minimum of four consecutive ribonucleotides, while type 2 enzymes require only one ribonucleotide for cleavage to occur. We reported the first crystal structure of a type 2 RNase H in complex with a nucleic acid substrate. It contains an RNA-DNA junction whose RNA residue is recognized by specific contacts of the 2-OH group with protein backbone and a conserved tyrosine residue. The same tyrosine residue stacks with the ribose ring of the DNA residue of the junction. Perfect stacking is possible only in the absence of a 2-OH group and this leads to a preference for a deoxyribonucleotide. Another consequence of the stacking interaction is a deformation of the nucleic acid geometry at the junction allowing the distorted phosphate group of the junction to participate in the coordination of the catalytic metal ion at the active site. Type 1 RNases H1 use a carboxylate to coordinate this metal ion, so the participation of the phosphate group in metal binding is a unique feature of type 2 enzymes. The two metal ions observed at the active site have very similar position to those seen previously in type 1 RNases H and other enzymes from retroviral integrase superfamily. One is responsible for the activation of the attacking nucleophile and the other for the stabilization of the transition state. Our structures provide the first structural insights into the specific cleavage of RNA-DNA junction, which may play an important role in DNA repair.
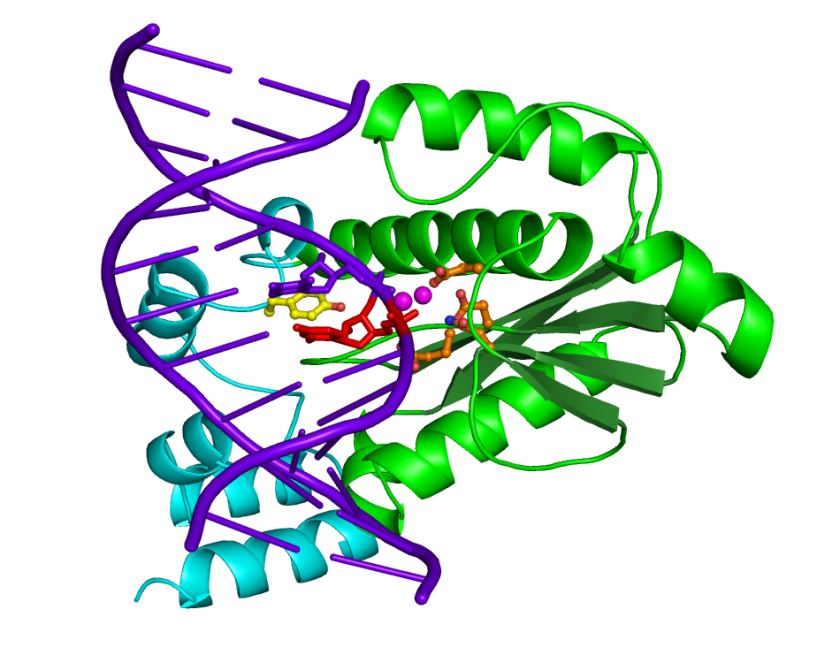
Figure 1. The overall structure of T. maritima RNase H2 in complex with nucleic acid substrate a 12-mer double-stranded DNA with single ribonucleotide. The protein is shown in green (the catalytic domain) and cyan (C-terminal helical domain). The nucleic acid is shown in purple with the single ribonucleotide in red. The carboxylate residues forming the active site are shown in orange ball-and-stick and the tyrosine residue at the heart of the RNA-DNA junction recognition module in yellow. Two magnesium ion bound at the active site are shown as purple spheres.