Joint Research Group Macromolecular Crystallography
Structure of the month - June 2004
Nat. Struct. Mol. Biol. Vol. 11, 2004, Pages 656-663
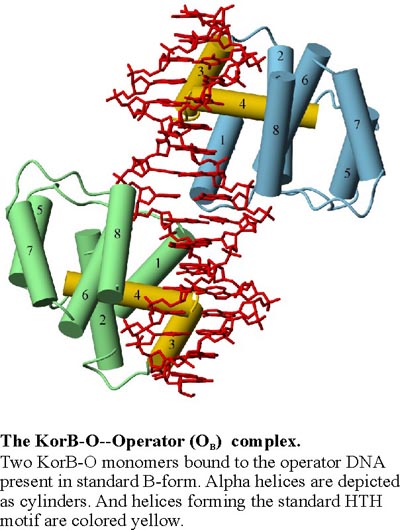
Crystal structure of ParB homolog KorB in complex with the operator DNA
Dheeraj Khare1, Günter Ziegelin2, Erich Lanka2 and Udo Heinemann1,3
1 Forschungsgruppe Kristallographie, Max-Delbrück-Centrum für Molekulare Medizin Berlin, Robert-Roessle-Str.10, 13125-Berlin, Germany
2 Max-Planck-Institut für Molekulare Genetik, Ihnestr. 73, D-14195 Berlin, Germany
3 Forschungsgruppe Kristallographie, Freie Universität Berlin, Takustr.6, 14195-Berlin, Germany
Abstract
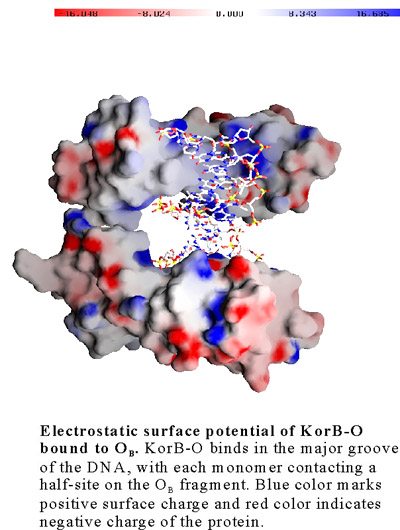
Species survival requires stringent control on how the genetic information is inherited to the offspring. Low-copy number genomes like bacterial chromosomes and certain plasmids have evolved partitioning (Par) mechanisms to ensure that daughter cells receive a full complement of the genetic material, which is in contrast to high-copy number plasmids that rely on random partitioning. All chromosomal or plasmid partitioning systems require three components, ParA a putative ATPase, ParB a sequence specific DNA binding protein and ParS a centromere like site. The KorB protein of broad host range plasmid RP4 plays a role in transcriptional regulation and also functions as a ParB homolog of plasmid's partitioning system. We have determined the X-ray structure of the DNA binding domain of KorB (KorB-O) in complex with a 17 bp DNA fragment containing the consensus OB sequence 5´-TTTAGCG/CGCTAAA-3´.
| Inflection Point | High energy remote | ||
| Diffraction experiment | |||
|---|---|---|---|
| 0.92022 | 0.92039 | 0.89844 | |
| Resolution range (Å) | 30-2.2 | 30-2.5 | 30-2.5 |
| Completeness (%) | 97.4 | 96.8 | 99.1 |
| < I / sigma(I)> | 14.9 | 12.3 | 20.7 |
| Rsym (%) | 7.7 | 12.3 | 6.1 |
| Refinement | |||
| Rwork (%) | 19.5 | ||
Rfree (%) | 25.0 | ||
Table 1. Crystallographic data from data collection at PSF-BL1 and refinement