Joint Research Group Macromolecular Crystallography
Structure of the month - June 2014
Nat. Struct. Mol. Biol. (2014) 389-396.
Ty3 reverse transcriptase complexed with an RNA-DNA hybrid shows structural and functional asymmetry
Elżbieta Nowaka, Jennifer T. Millerb, Marion K. Bonab, Justyna Studnickaa, Roman H. Szczepanowskic, Jakub Jurkowskia, Stuart F. J. Le Griceb*, Marcin Nowotnya*
a Laboratory of Protein Structure, Internal Institute of Molecular and Cell Biology, Warsaw, Poland
b Reverse Transcriptase Biochemistry Section, HIV Drug Resistance Program, Frederick, MD, USA
c Biophysics Core Facility, Internal Institute of Molecular and Cell Biology, Warsaw, Poland
* Corresponding author 1: Stuart F. J. Le Grice, Reverse Transcriptase Biochemistry Section, HIV Drug Resistance Program, Frederick National Laboratory, Frederick, MD, USA
E-mail: legrices@mail.nih.gov
Phone: +1-301-846-5256
* Corresponding author 2: Marcin Nowotny, Laboratory of Protein Structure, Internal Institute of Molecular and Cell Biology, Warsaw, Poland
E-mail: mnowotny@iimcb.gov.pl
Phone: +48(0)22-597-0717
Abstract
Retrotransposons are mobile genetic elements that replicate through an RNA intermediate. They are one of the most potent forces shaping eukaryotic genomes with around 40% of the human genome derived from them. A critical step in their proliferation is reverse transcription in which single-stranded RNA is converted to double-stranded DNA. This process is catalyzed by an enzyme called reverse transcriptase (RT) which has two activities: DNA polymerase for DNA synthesis and RNase H for the degradation of the RNA in the intermediate RNA/DNA hybrid.
We solved the first crystal structure of a retrotransposon RT – that of a complex of yeast element Ty3 RT in complex with an RNA/DNA substrate. It revealed an unanticipated homodimeric architecture of the enzyme. One of the subunits has a conformation similar to other RT polymerases, while the other has an altered conformation with the polymerase active site blocked. Our structural and biochemical data showed that the two activities: polymerase and RNase H, are located in two separate subunits of the homodimer. We also postulated that the RNase H domain undergoes a large conformational change in order to cleave the substrate, which serves to regulate its activity.
Retroviruses evolved from retrotransposons and the best-characterized viral RT is from the human immunodeficiency virus (HIV). It is a heterodimer of larger subunit and a smaller one that lacks the RNase H domain. The overall architecture of HIV and Ty3 RTs is similar but several crucial differences are observed. Ty3 RT is a substrate-induced homodimer, while HIV RT is a constitutive heterodimer. Moreover, in HIV RT both activities reside in the larger subunit and in Ty3 enzyme they are located in different subunits of the homodimer. We provided the first structural and mechanistic information for a retrotransposon RT with implications for the evolution of retroviruses.
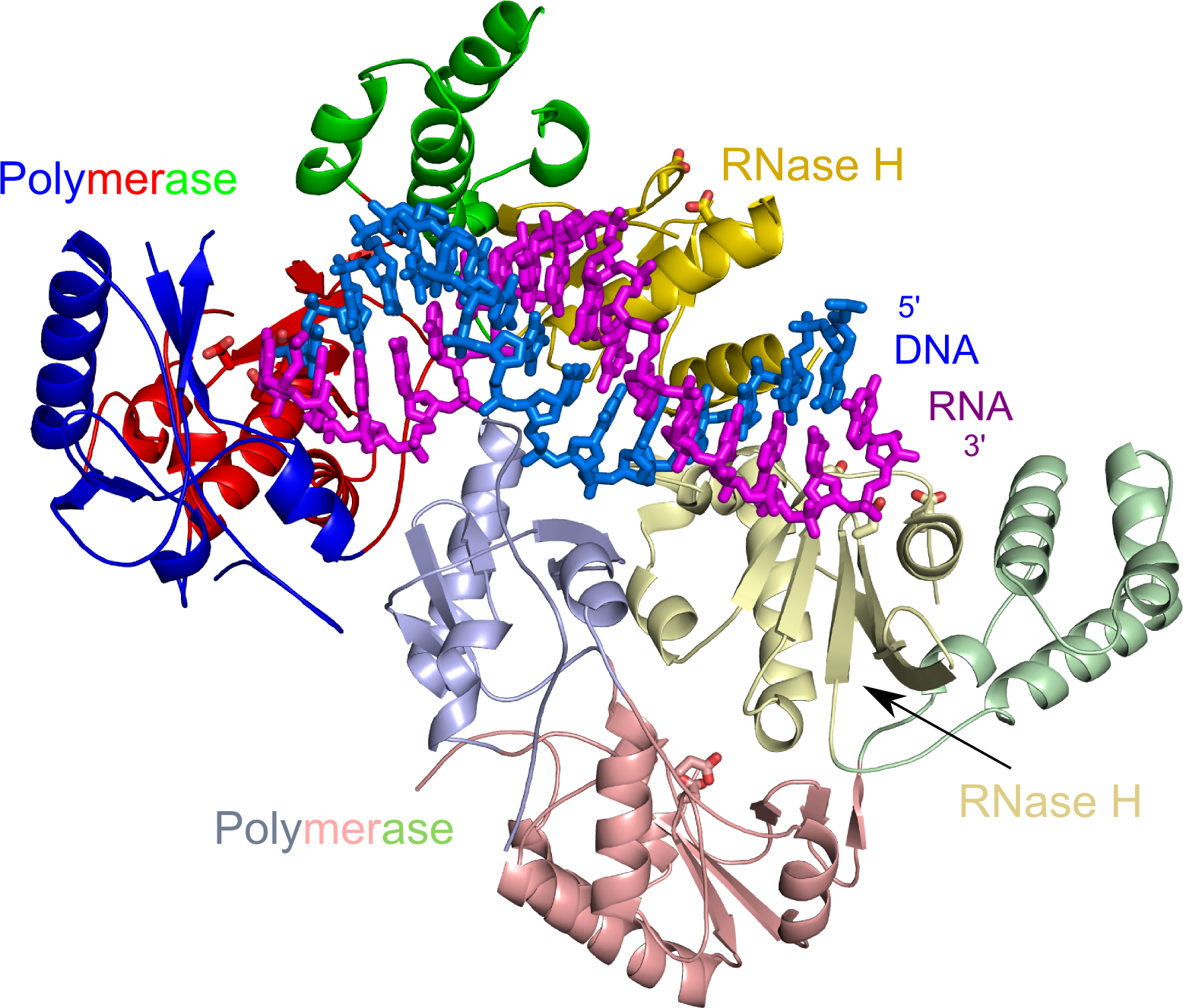
Overall structure of Ty3 RT in complex with an RNA/DNA hybrid. The polymerase subdomains are colored blue, red and green and the RNase H domains in yellow. Lighter shades of the colors are used for the subunit with altered conformation and blocked polymerase active site. The RNA strand of the hybrid is in purple and the DNA strand in marine. Active site residues are shown as sticks.

