Joint Research Group Macromolecular Crystallography
Structure of the month - January 2008
Science Vol. 319, 2008, Pages 206 - 209
Designed Protein-Protein Association
Dirk Grueninger, Nora Treiber, Mathias O. P. Ziegler, Jochen W. A. Koetter, Monika-Sarah Schulze, Georg E. Schulz*
Institute for Organic Chemistry and Biochemistry, Albert-Ludwigs-University, Albertstrasse 21, 79104 Freiburg im Breisgau, Germany
*To whom correspondence should be addressed. E-mail: georg.schulz@ocbc.uni-freiburg.de
Abstract
Protein association is a central feature of all living organisms. Numerous natural contact interfaces have been established and analyzed at atomic resolution, yielding some rather general rules governing the association between protein subunits. We have applied these rules to produce a number of novel assemblies, pointing out that a given protein can be engineered to form contacts at various points of its surface. Our results demonstrate that symmetry plays an important role because it determines the multiplicity of a designed contact and therefore the number of required mutations. Several proteins required only a single side-chain modification to associate to a higher-order complex. The mobility of the side chains which become buried into the newly designed contact has also to be taken into account. The results of our efforts were checked by gelpermeation and dynamic light scattering. Four assemblies have been structurally elucidated. Furthermore the combination of different contact regions yielded in fiber associations. Comparisons between the designed and the obtained contacts provide useful guidelines for the development of future architectures.
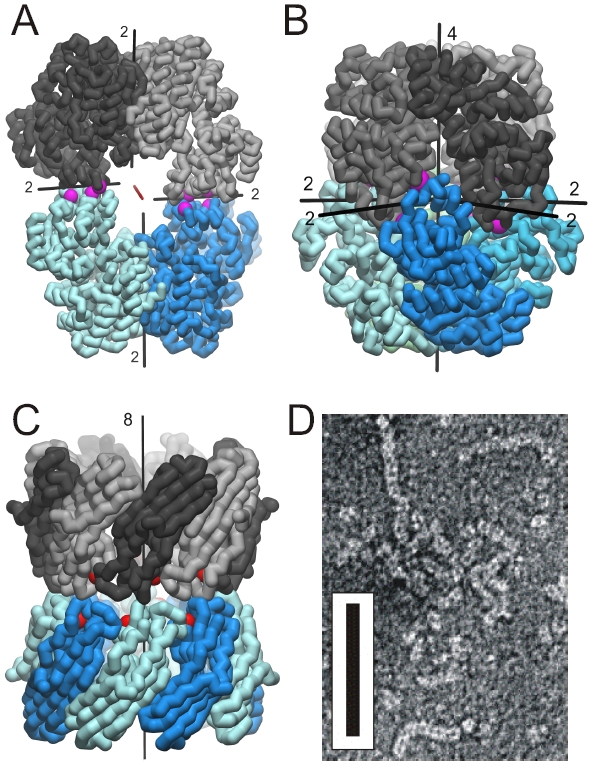
Figure 1. Established oligomer structures. The mutations were introduced to produce symmetric associations and are marked by purple spheres. (A) Crystal structure of C2-symmetric urocanase (Uro) showing the twofold molecular symmetry axis (red) and four local twofold axes. (B) D4-symmetric octameric association of L-Rhamnulose-1- phosphate aldolase (Rua). (C) D8-symmetric association of two rim-domain fragments of mycobacterial porin (Myp). The positions of the 52-residue deletions are marked by red spheres. (D) Negatively stained electron micrograph of multimeric Rua showing the fiber association yielded by combination of two obtained interfaces. The scale is given by a black bar with the dimension of 100 nm x 10 nm.